|
1 | 1 | # cotengrust |
2 | 2 |
|
3 | | -`cotengrust` provides fast rust implemented versions of contraction ordering |
4 | | -primitives for tensor networks and einsums. |
| 3 | +`cotengrust` provides fast rust implementations of contraction ordering |
| 4 | +primitives for tensor networks or einsum expressions. The two main functions |
| 5 | +are: |
| 6 | + |
| 7 | +- `optimize_optimal(inputs, output, size_dict, **kwargs)` |
| 8 | +- `optimize_greedy(inputs, output, size_dict, **kwargs)` |
| 9 | + |
| 10 | +The optimal algorithm is an optimized version of the `opt_einsum` 'dp' |
| 11 | +path - itself an implementation of https://arxiv.org/abs/1304.6112. |
| 12 | + |
| 13 | + |
| 14 | +## Installation |
| 15 | + |
| 16 | +`cotengrust` is available for most platforms from |
| 17 | +[PyPI](https://pypi.org/project/cotengrust/): |
| 18 | + |
| 19 | +```bash |
| 20 | +pip install cotengrust |
| 21 | +``` |
| 22 | + |
| 23 | +or if you want to develop locally (which requires [pyo3](https://github.com/PyO3/pyo3) |
| 24 | +and [maturin](https://github.com/PyO3/maturin)): |
| 25 | + |
| 26 | +```bash |
| 27 | +git clone https://github.com/jcmgray/cotengrust.git |
| 28 | +cd cotengrust |
| 29 | +maturin develop --release |
| 30 | +``` |
| 31 | +(the release flag is very important for assessing performance!). |
| 32 | + |
| 33 | + |
| 34 | +## Usage |
| 35 | + |
| 36 | +If `cotengrust` is installed, then by default `cotengra` will use it for its |
| 37 | +greedy and optimal subroutines, notably subtree reconfiguration. You can also |
| 38 | +call the routines directly: |
| 39 | + |
| 40 | +```python |
| 41 | +import cotengra as ctg |
| 42 | +import cotengrust as ctgr |
| 43 | + |
| 44 | +# specify an 8x8 square lattice contraction |
| 45 | +inputs, output, shapes, size_dict = ctg.utils.lattice_equation([8, 8]) |
| 46 | + |
| 47 | +# find the optimal 'combo' contraction path |
| 48 | +%%time |
| 49 | +path = ctgr.optimize_optimal(inputs, output, size_dict, minimize='combo') |
| 50 | +# CPU times: user 13.7 s, sys: 83.4 ms, total: 13.7 s |
| 51 | +# Wall time: 13.7 s |
| 52 | + |
| 53 | +# construct a contraction tree for further introspection |
| 54 | +tree = ctg.ContractionTree.from_path( |
| 55 | + inputs, output, size_dict, path=path |
| 56 | +) |
| 57 | +tree.plot_rubberband() |
| 58 | +``` |
| 59 | +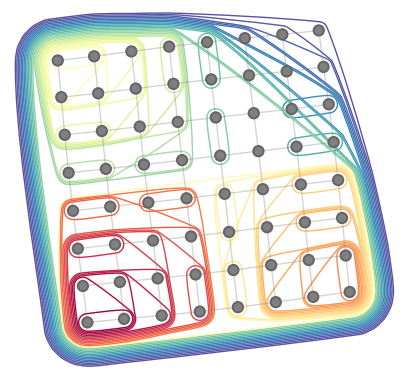 |
| 60 | + |
| 61 | + |
| 62 | +## API |
| 63 | + |
| 64 | +The optimize functions follow the api of the python implementations in `cotengra.pathfinders.path_basic.py`. |
| 65 | + |
| 66 | +```python |
| 67 | +def optimize_optimal( |
| 68 | + inputs, |
| 69 | + output, |
| 70 | + size_dict, |
| 71 | + minimize='flops', |
| 72 | + cost_cap=2, |
| 73 | + search_outer=False, |
| 74 | + simplify=True, |
| 75 | + use_ssa=False, |
| 76 | +): |
| 77 | + """Find an optimal contraction ordering. |
| 78 | +
|
| 79 | + Parameters |
| 80 | + ---------- |
| 81 | + inputs : Sequence[Sequence[str]] |
| 82 | + The indices of each input tensor. |
| 83 | + output : Sequence[str] |
| 84 | + The indices of the output tensor. |
| 85 | + size_dict : dict[str, int] |
| 86 | + The size of each index. |
| 87 | + minimize : str, optional |
| 88 | + The cost function to minimize. The options are: |
| 89 | +
|
| 90 | + - "flops": minimize with respect to total operation count only |
| 91 | + (also known as contraction cost) |
| 92 | + - "size": minimize with respect to maximum intermediate size only |
| 93 | + (also known as contraction width) |
| 94 | + - 'write' : minimize the sum of all tensor sizes, i.e. memory written |
| 95 | + - 'combo' or 'combo={factor}` : minimize the sum of |
| 96 | + FLOPS + factor * WRITE, with a default factor of 64. |
| 97 | + - 'limit' or 'limit={factor}` : minimize the sum of |
| 98 | + MAX(FLOPS, alpha * WRITE) for each individual contraction, with a |
| 99 | + default factor of 64. |
| 100 | +
|
| 101 | + 'combo' is generally a good default in term of practical hardware |
| 102 | + performance, where both memory bandwidth and compute are limited. |
| 103 | + cost_cap : float, optional |
| 104 | + The maximum cost of a contraction to initially consider. This acts like |
| 105 | + a sieve and is doubled at each iteration until the optimal path can |
| 106 | + be found, but supplying an accurate guess can speed up the algorithm. |
| 107 | + search_outer : bool, optional |
| 108 | + If True, consider outer product contractions. This is much slower but |
| 109 | + theoretically might be required to find the true optimal 'flops' |
| 110 | + ordering. In practical settings (i.e. with minimize='combo'), outer |
| 111 | + products should not be required. |
| 112 | + simplify : bool, optional |
| 113 | + Whether to perform simplifications before optimizing. These are: |
| 114 | +
|
| 115 | + - ignore any indices that appear in all terms |
| 116 | + - combine any repeated indices within a single term |
| 117 | + - reduce any non-output indices that only appear on a single term |
| 118 | + - combine any scalar terms |
| 119 | + - combine any tensors with matching indices (hadamard products) |
| 120 | +
|
| 121 | + Such simpifications may be required in the general case for the proper |
| 122 | + functioning of the core optimization, but may be skipped if the input |
| 123 | + indices are already in a simplified form. |
| 124 | + use_ssa : bool, optional |
| 125 | + Whether to return the contraction path in 'single static assignment' |
| 126 | + (SSA) format (i.e. as if each intermediate is appended to the list of |
| 127 | + inputs, without removals). This can be quicker and easier to work with |
| 128 | + than the 'linear recycled' format that `numpy` and `opt_einsum` use. |
| 129 | +
|
| 130 | + Returns |
| 131 | + ------- |
| 132 | + path : list[list[int]] |
| 133 | + The contraction path, given as a sequence of pairs of node indices. It |
| 134 | + may also have single term contractions if `simplify=True`. |
| 135 | + """ |
| 136 | + ... |
| 137 | + |
| 138 | + |
| 139 | +def optimize_greedy( |
| 140 | + inputs, |
| 141 | + output, |
| 142 | + size_dict, |
| 143 | + costmod=1.0, |
| 144 | + temperature=0.0, |
| 145 | + simplify=True, |
| 146 | + use_ssa=False, |
| 147 | +): |
| 148 | + """Find a contraction path using a (randomizable) greedy algorithm. |
| 149 | +
|
| 150 | + Parameters |
| 151 | + ---------- |
| 152 | + inputs : Sequence[Sequence[str]] |
| 153 | + The indices of each input tensor. |
| 154 | + output : Sequence[str] |
| 155 | + The indices of the output tensor. |
| 156 | + size_dict : dict[str, int] |
| 157 | + A dictionary mapping indices to their dimension. |
| 158 | + costmod : float, optional |
| 159 | + When assessing local greedy scores how much to weight the size of the |
| 160 | + tensors removed compared to the size of the tensor added:: |
| 161 | +
|
| 162 | + score = size_ab - costmod * (size_a + size_b) |
| 163 | +
|
| 164 | + This can be a useful hyper-parameter to tune. |
| 165 | + temperature : float, optional |
| 166 | + When asessing local greedy scores, how much to randomly perturb the |
| 167 | + score. This is implemented as:: |
| 168 | +
|
| 169 | + score -> sign(score) * log(|score|) - temperature * gumbel() |
| 170 | +
|
| 171 | + which implements boltzmann sampling. |
| 172 | + simplify : bool, optional |
| 173 | + Whether to perform simplifications before optimizing. These are: |
| 174 | +
|
| 175 | + - ignore any indices that appear in all terms |
| 176 | + - combine any repeated indices within a single term |
| 177 | + - reduce any non-output indices that only appear on a single term |
| 178 | + - combine any scalar terms |
| 179 | + - combine any tensors with matching indices (hadamard products) |
| 180 | +
|
| 181 | + Such simpifications may be required in the general case for the proper |
| 182 | + functioning of the core optimization, but may be skipped if the input |
| 183 | + indices are already in a simplified form. |
| 184 | + use_ssa : bool, optional |
| 185 | + Whether to return the contraction path in 'single static assignment' |
| 186 | + (SSA) format (i.e. as if each intermediate is appended to the list of |
| 187 | + inputs, without removals). This can be quicker and easier to work with |
| 188 | + than the 'linear recycled' format that `numpy` and `opt_einsum` use. |
| 189 | +
|
| 190 | + Returns |
| 191 | + ------- |
| 192 | + path : list[list[int]] |
| 193 | + The contraction path, given as a sequence of pairs of node indices. It |
| 194 | + may also have single term contractions if `simplify=True`. |
| 195 | + """ |
| 196 | + |
| 197 | +def optimize_simplify( |
| 198 | + inputs, |
| 199 | + output, |
| 200 | + size_dict, |
| 201 | + use_ssa=False, |
| 202 | +): |
| 203 | + """Find the (partial) contracton path for simplifiactions only. |
| 204 | +
|
| 205 | + Parameters |
| 206 | + ---------- |
| 207 | + inputs : Sequence[Sequence[str]] |
| 208 | + The indices of each input tensor. |
| 209 | + output : Sequence[str] |
| 210 | + The indices of the output tensor. |
| 211 | + size_dict : dict[str, int] |
| 212 | + A dictionary mapping indices to their dimension. |
| 213 | + use_ssa : bool, optional |
| 214 | + Whether to return the contraction path in 'single static assignment' |
| 215 | + (SSA) format (i.e. as if each intermediate is appended to the list of |
| 216 | + inputs, without removals). This can be quicker and easier to work with |
| 217 | + than the 'linear recycled' format that `numpy` and `opt_einsum` use. |
| 218 | +
|
| 219 | + Returns |
| 220 | + ------- |
| 221 | + path : list[list[int]] |
| 222 | + The contraction path, given as a sequence of pairs of node indices. It |
| 223 | + may also have single term contractions. |
| 224 | +
|
| 225 | + """ |
| 226 | + ... |
| 227 | + |
| 228 | +def ssa_to_linear(ssa_path, n=None): |
| 229 | + """Convert a SSA path to linear format.""" |
| 230 | + ... |
| 231 | + |
| 232 | +def find_subgraphs(inputs, output, size_dict,): |
| 233 | + """Find all disconnected subgraphs of a specified contraction.""" |
| 234 | + ... |
| 235 | +``` |
| 236 | + |
0 commit comments